|
|
Botella-Soler, V., Valderrama, M., Crepon, B., Navarro, V., & Le Van Quyen, M. (2012). Large-Scale Cortical Dynamics of Sleep Slow Waves. PLoS One, 7(2), e30757–10pp.

Abstract: Slow waves constitute the main signature of sleep in the electroencephalogram (EEG). They reflect alternating periods of neuronal hyperpolarization and depolarization in cortical networks. While recent findings have demonstrated their functional role in shaping and strengthening neuronal networks, a large-scale characterization of these two processes remains elusive in the human brain. In this study, by using simultaneous scalp EEG and intracranial recordings in 10 epileptic subjects, we examined the dynamics of hyperpolarization and depolarization waves over a large extent of the human cortex. We report that both hyperpolarization and depolarization processes can occur with two different characteristic time durations which are consistent across all subjects. For both hyperpolarization and depolarization waves, their average speed over the cortex was estimated to be approximately 1 m/s. Finally, we characterized their propagation pathways by studying the preferential trajectories between most involved intracranial contacts. For both waves, although single events could begin in almost all investigated sites across the entire cortex, we found that the majority of the preferential starting locations were located in frontal regions of the brain while they had a tendency to end in posterior and temporal regions.
|
|
|
|
Sanjuan, R., Nebot, M., Peris, J. B., & Alcami, J. (2013). Immune Activation Promotes Evolutionary Conservation of T-Cell Epitopes in HIV-1. PLoS. Biol., 11(4), e1001523–10pp.

Abstract: The immune system should constitute a strong selective pressure promoting viral genetic diversity and evolution. However, HIV shows lower sequence variability at T-cell epitopes than elsewhere in the genome, in contrast with other human RNA viruses. Here, we propose that epitope conservation is a consequence of the particular interactions established between HIV and the immune system. On one hand, epitope recognition triggers an anti-HIV response mediated by cytotoxic T-lymphocytes (CTLs), but on the other hand, activation of CD4(+) helper T lymphocytes (T-H cells) promotes HIV replication. Mathematical modeling of these opposite selective forces revealed that selection at the intrapatient level can promote either T-cell epitope conservation or escape. We predict greater conservation for epitopes contributing significantly to total immune activation levels (immunodominance), and when T-H cell infection is concomitant to epitope recognition (transinfection). We suggest that HIV-driven immune activation in the lymph nodes during the chronic stage of the disease may offer a favorable scenario for epitope conservation. Our results also support the view that some pathogens draw benefits from the immune response and suggest that vaccination strategies based on conserved T-H epitopes may be counterproductive.
|
|
|
|
n_TOF Collaboration(Gawlik, A. et al), Domingo-Pardo, C., Tain, J. L., & Tarifeño-Saldivia, A. (2021). Radiative Neutron Capture Cross-Section Measurement of Ge Isotopes at n_TOF CERN Facility and Its Importance for Stellar Nucleosynthesis. Acta Phys. Pol. A, 139(4), 383–388.
Abstract: This manuscript summarizes the results of radiative neutron capture cross-section measurements on two stable germanium isotopes, Ge-70 and Ge-73. Experiments were performed at the n_TOF facility at CERN via the time-of-flight technique, over a wide neutron energy range, for all stable germanium isotopes (70,72,73,74, and 76). Results for Ge-70 [Phys. Rev. C 100, 045804 (2019)] and Ge-73 [Phys. Lett. B 790, 458 (2019)] are already published. In the field of nuclear structure, such measurements allow to study excited levels close to the neutron binding energy and to obtain information on nuclear properties. In stellar nucleosynthesis research, neutron induced reactions on germanium are of importance for nucleosynthesis in the weak component of the slow neutron capture processes.
|
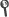
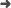
|
|
|
Albiol, A., Albiol, F., Paredes, R., Plasencia-Martinez, J. M., Blanco Barrio, A., Garcia Santos, J. M., et al. (2022). A comparison of Covid-19 early detection between convolutional neural networks and radiologists. Insights Imaging, 13(1), 122–12pp.

Abstract: Background The role of chest radiography in COVID-19 disease has changed since the beginning of the pandemic from a diagnostic tool when microbiological resources were scarce to a different one focused on detecting and monitoring COVID-19 lung involvement. Using chest radiographs, early detection of the disease is still helpful in resource-poor environments. However, the sensitivity of a chest radiograph for diagnosing COVID-19 is modest, even for expert radiologists. In this paper, the performance of a deep learning algorithm on the first clinical encounter is evaluated and compared with a group of radiologists with different years of experience. Methods The algorithm uses an ensemble of four deep convolutional networks, Ensemble4Covid, trained to detect COVID-19 on frontal chest radiographs. The algorithm was tested using images from the first clinical encounter of positive and negative cases. Its performance was compared with five radiologists on a smaller test subset of patients. The algorithm's performance was also validated using the public dataset COVIDx. Results Compared to the consensus of five radiologists, the Ensemble4Covid model achieved an AUC of 0.85, whereas the radiologists achieved an AUC of 0.71. Compared with other state-of-the-art models, the performance of a single model of our ensemble achieved nonsignificant differences in the public dataset COVIDx. Conclusion The results show that the use of images from the first clinical encounter significantly drops the detection performance of COVID-19. The performance of our Ensemble4Covid under these challenging conditions is considerably higher compared to a consensus of five radiologists. Artificial intelligence can be used for the fast diagnosis of COVID-19.
|
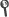
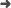
|
|
|
Fernandez Casani, A., Garcia Montoro, C., Gonzalez de la Hoz, S., Salt, J., Sanchez, J., & Villaplana Perez, M. (2023). Big Data Analytics for the ATLAS EventIndex Project with Apache Spark. Comput. Math. Methods, 2023, 6900908–19pp.

Abstract: The ATLAS EventIndex was designed to provide a global event catalogue and limited event-level metadata for ATLAS experiment of the Large Hadron Collider (LHC) and their analysis groups and users during Run 2 (2015-2018) and has been running in production since. The LHC Run 3, started in 2022, has seen increased data-taking and simulation production rates, with which the current infrastructure would still cope but may be stretched to its limits by the end of Run 3. A new core storage service is being developed in HBase/Phoenix, and there is work in progress to provide at least the same functionality as the current one for increased data ingestion and search rates and with increasing volumes of stored data. In addition, new tools are being developed for solving the needed access cases within the new storage. This paper describes a new tool using Spark and implemented in Scala for accessing the big data quantities of the EventIndex project stored in HBase/Phoenix. With this tool, we can offer data discovery capabilities at different granularities, providing Spark Dataframes that can be used or refined within the same framework. Data analytic cases of the EventIndex project are implemented, like the search for duplicates of events from the same or different datasets. An algorithm and implementation for the calculation of overlap matrices of events across different datasets are presented. Our approach can be used by other higher-level tools and users, to ease access to the data in a performant and standard way using Spark abstractions. The provided tools decouple data access from the actual data schema, which makes it convenient to hide complexity and possible changes on the backed storage.
|
|

