|
|
Cervantes, D., Fioresi, R., Lledo, M. A., & Nadal, F. A. (2012). Quadratic deformation of Minkowski space. Fortschritte Phys.-Prog. Phys., 60(9-10), 970–976.
Abstract: We present a deformation of the Minkowski space as embedded into the conformal space (in the formalism of twistors) based in the quantum versions of the corresponding kinematic groups. We compute explicitly the star product, whose Poisson bracket is quadratic. We show that the star product although defined on the polynomials can be extended differentiably. Finally we compute the Eucliden and Minkowskian real forms of the deformation.
|
|
|
|
Galli, P., Ortin, T., Perz, J., & Shahbazi, C. S. (2012). From supersymmetric to non-supersymmetric black holes. Fortschritte Phys.-Prog. Phys., 60(9-10), 1026–1029.
Abstract: Methods similar to those used for obtaining supersymmetric black hole solutions can be employed to find also non-supersymmetric solutions. We briefly review some of them, with the emphasis on the non-extremal deformation ansatz of [1].
|
|
|
|
Tortola, M. (2013). Status of three-neutrino oscillation parameters. Fortschritte Phys.-Prog. Phys., 61(4-5), 427–440.
Abstract: Here we review the current status of global fits to neutrino oscillation data within the three-flavour framework. In our analysis we include the most recent data from solar and atmospheric neutrino experiments as well as the latest results from the long-baseline accelerator neutrino experiments and the recent measurements of reactor neutrino disappearance reported by Double Chooz, Daya Bay and RENO. We present updated determinations for the two neutrino mass splittings and the three mixing angles responsible for neutrino oscillations that, for the first time, have all been measured with 1 sigma accuracies ranging from 3 to 15%. A weak sensitivity for the CP violating phase is also reported from the global analysis.
|
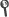
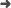
|
|
|
Morisi, S., & Valle, J. W. F. (2013). Neutrino masses and mixing: a flavour symmetry roadmap. Fortschritte Phys.-Prog. Phys., 61(4-5), 466–492.
Abstract: Over the last ten years tri-bimaximal mixing has played an important role in modeling the flavour problem. We give a short review of the status of flavour symmetry models of neutrino mixing. We concentrate on non-Abelian discrete symmetries, which provide a simple way to account for the TBM pattern. We discuss phenomenological implications such as neutrinoless double beta decay, lepton flavour violation as well as theoretical aspects such as the possibility to explain quarks and leptons within a common framework, such as grand unified models.
|
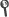
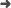
|
|
|
Peris, J. B., Davis, P., Cuevas, J. M., Nebot, M., & Sanjuan, R. (2010). Distribution of Fitness Effects Caused by Single-Nucleotide Substitutions in Bacteriophage f1. Genetics, 185(2), 603–U308.

Abstract: Empirical knowledge of the fitness effects of mutations is important for understanding many evolutionary processes, yet this knowledge is often hampered by several sources of measurement error and bias. Most of these problems can be solved using site-directed mutagenesis to engineer single mutations, an approach particularly suited for viruses due to their small genomes. Here, we used this technique to measure the fitness effect of 100 single-nucleotide substitutions in the bacteriophage f1, a filamentous single-strand DNA virus. We found that approximately one-fifth of all mutations are lethal. Viable ones reduced fitness by 11% on average and were accurately described by a log-normal distribution. More than 90% of synonymous substitutions were selectively neutral, while those affecting intergenic regions reduced fitness by 14% on average. Mutations leading to amino acid substitutions had an overall mean deleterious effect of 37%, which increased to 45% for those changing the amino acid polarity. Interestingly, mutations affecting early steps of the infection cycle tended to be more deleterious than those affecting late steps. Finally, we observed at least two beneficial mutations. Our results confirm that high mutational sensitivity is a general property of viruses with small genomes, including RNA and single-strand DNA viruses infecting animals, plants, and bacteria.
|
|

