|
|
LHCb Collaboration(Aaij, R. et al), Martinez-Vidal, F., Oyanguren, A., Ruiz Valls, P., & Sanchez Mayordomo, C. (2015). Measurement of the track reconstruction efficiency at LHCb. J. Instrum., 10, P02007–23pp.
Abstract: The determination of track reconstruction efficiencies at LHCb using J/psi -> mu(+)mu(-) decays is presented. Efficiencies above 95% are found for the data taking periods in 2010, 2011, and 2012. The ratio of the track reconstruction efficiency of muons in data and simulation is compatible with unity and measured with an uncertainty of 0.8% for data taking in 2010, and at a precision of 0.4% for data taking in 2011 and 2012. For hadrons an additional 1.4% uncertainty due to material interactions is assumed. This result is crucial for accurate cross section and branching fraction measurements in LHCb.
|
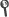
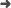
|
|
|
Ortega, P. G., Torres-Espallardo, I., Cerutti, F., Ferrari, A., Gillam, J. E., Lacasta, C., et al. (2015). Noise evaluation of Compton camera imaging for proton therapy. Phys. Med. Biol., 60(5), 1845–1863.

Abstract: Compton Cameras emerged as an alternative for real-time dose monitoring techniques for Particle Therapy (PT), based on the detection of prompt-gammas. As a consequence of the Compton scattering process, the gamma origin point can be restricted onto the surface of a cone (Compton cone). Through image reconstruction techniques, the distribution of the gamma emitters can be estimated, using cone-surfaces backprojections of the Compton cones through the image space, along with more sophisticated statistical methods to improve the image quality. To calculate the Compton cone required for image reconstruction, either two interactions, the last being photoelectric absorption, or three scatter interactions are needed. Because of the high energy of the photons in PT the first option might not be adequate, as the photon is not absorbed in general. However, the second option is less efficient. That is the reason to resort to spectral reconstructions, where the incoming. energy is considered as a variable in the reconstruction inverse problem. Jointly with prompt gamma, secondary neutrons and scattered photons, not strongly correlated with the dose map, can also reach the imaging detector and produce false events. These events deteriorate the image quality. Also, high intensity beams can produce particle accumulation in the camera, which lead to an increase of random coincidences, meaning events which gather measurements from different incoming particles. The noise scenario is expected to be different if double or triple events are used, and consequently, the reconstructed images can be affected differently by spurious data. The aim of the present work is to study the effect of false events in the reconstructed image, evaluating their impact in the determination of the beam particle ranges. A simulation study that includes misidentified events (neutrons and random coincidences) in the final image of a Compton Telescope for PT monitoring is presented. The complete chain of detection, from the beam particle entering a phantom to the event classification, is simulated using FLUKA. The range determination is later estimated from the reconstructed image obtained from a two and three-event algorithm based on Maximum Likelihood Expectation Maximization. The neutron background and random coincidences due to a therapeutic-like time structure are analyzed for mono-energetic proton beams. The time structure of the beam is included in the simulations, which will affect the rate of particles entering the detector.
|
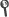
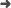
|
|
|
Egea Canet, F. J. et al, Gadea, A., & Huyuk, T. (2015). Digital Front-End Electronics for the Neutron Detector NEDA. IEEE Trans. Nucl. Sci., 62(3), 1063–1069.
Abstract: This paper presents the design of the NEDA (Neutron Detector Array) electronics, a first attempt to involve the use of digital electronics in large neutron detector arrays. Starting from the front-end modules attached to the PMTs (PhotoMultiplier Tubes) and ending up with the data processing workstations, a comprehensive electronic system capable of dealing with the acquisition and pre-processing of the neutron array is detailed. Among the electronic modules required, we emphasize the front-end analog processing, the digitalization, digital pre-processing and communications firmware, as well as the integration of the GTS (Global Trigger and Synchronization) system, already used successfully in AGATA (Advanced Gamma Tracking Array). The NEDA array will be available for measurements in 2016.
|
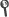
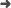
|
|
|
Egea Canet, F. J. et al, Gadea, A., & Huyuk, T. (2015). A New Front-End High-Resolution Sampling Board for the New-Generation Electronics of EXOGAM2 and NEDA Detectors. IEEE Trans. Nucl. Sci., 62(3), 1056–1062.
Abstract: This paper presents the final design and results of the FADC Mezzanine for the EXOGAM (EXOtic GAMma array spectrometer) and NEDA (Neutron Detector Array) detectors. The measurements performed include those of studying the effective number of bits, the energy resolution using HP-Ge detectors, as well as timing histograms and discrimination performance. Finally, the conclusion shows how a common digitizing device has been integrated in the experimental environment of two very different detectors which combine both low-noise acquisition and fast sampling rates. Not only the integration fulfilled the expected specifications on both systems, but it also showed how a study of synergy between detectors could lead to the reduction of resources and time by applying a common strategy.
|
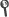
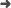
|
|
|
Ames, S. K., Gardner, S. N., Marti, J. M., Slezak, T. R., Gokhale, M. B., & Allen, J. E. (2015). Using populations of human and microbial genomes for organism detection in metagenomes. Genome Res., 25(7), 1056–1067.

Abstract: Identifying causative disease agents in human patients from shotgun metagenomic sequencing (SMS) presents a powerful tool to apply when other targeted diagnostics fail. Numerous technical challenges remain, however, before SMS can move beyond the role of research tool. Accurately separating the known and unknown organism content remains difficult, particularly when SMS is applied as a last resort. The true amount of human DNA that remains in a sample after screening against the human reference genome and filtering nonbiological components left from library preparation has previously been underreported. In this study, we create the most comprehensive collection of microbial and reference-free human genetic variation available in a database optimized for efficient metagenomic search by extracting sequences from GenBank and the 1000 Genomes Project. The results reveal new human sequences found in individual Human Microbiome Project (HMP) samples. Individual samples contain up to 95% human sequence, and 4% of the individual HMP samples contain 10% or more human reads. Left unidentified, human reads can complicate and slow down further analysis and lead to inaccurately labeled microbial taxa and ultimately lead to privacy concerns as more human genome data is collected.
|
|

