|
|
BABAR Collaboration(del Amo Sanchez, P. et al), Lopez-March, N., Martinez-Vidal, F., Milanes, D. A., & Oyanguren, A. (2010). Search for b -> u transitions in B- -> DK- and D*K- decays. Phys. Rev. D, 82(7), 072006–18pp.

Abstract: We report results from an updated study of the suppressed decays We report results from an updated study of the suppressed decays B- -> DK- and B- -> D*K- followed by D -> K+ pi(-), where D-(*()0) indicates a D-(*()0)or a (D) over bar (()*()0) meson, and D* -> D pi(0) or D* -> D gamma. These decays are sensitive to the Cabibbo-Kobayashi-Maskawa unitarity triangle angle gamma due to interference between the b -> c transition B- -> D-(*K-)0(-) followed by the doubly Cabibbo-suppressed decay D-0 -> K+ pi(-), and the b -> u transition B- -> (D) over bar (()*()0) K- followed by the Cabibbo-favored decay (D) over bar (0) -> K+ pi(-). We also report an analysis of the decay B- -> D-(*())pi(-) with the D decaying into the doubly Cabibbo-suppressed mode D -> K+ pi(-). Our results are based on 467 x 10(6) Gamma(4S) -> BB- decays collected with the BABAR detector at SLAC. We measure the ratios R-(*()) of the suppressed ([K+ pi(-)](D)K- / pi(-)) to favored ([K+ pi(-)](D)K- / pi(-)) branching fractions as well as the CP asymmetries A(()*()) of those modes. We see indications of signals for the B- -> DK- and B- -> D-D pi 0(()*()) K- suppressed modes, with statistical significances of 2.1 and 2.2 sigma, respectively, and we measure: R-DK = (1.1 +/- 0: 6 +/- 0.2) x 10(-2); A(DK) = -0.86 +/- 0: 47(-0.16)(+0.12), R-(D pi 0)K* = (1.8 +/- 0: 9 +/- 0: 4) x 10(-2); A ((D pi 0)K)* = +0.77 +/- 0: 35 +/- 0.12; R-(D gamma)K* = (1.3 +/- 1.4 +/- 0.8) x 10(-2); A((D gamma)K)* = +0.36 +/- 0: 94(-0.41)(+0.25), where the first uncertainty is statistical and the second is systematic. We use a frequentist approach to obtain the magnitude of the ratio r(B) equivalent to vertical bar A(B- -> (D) over bar 0K(-))/A(B- -> (DK-)-K-0)vertical bar = (9.5(-4.1)(+5.1))%, with r(B) < 16: 7% at 90% confidence level. In the case of B- -> D*K- we find r(B) equivalent to vertical bar A(B- -> <(D)over bar>0K(-))/A(B- -> (DK-)-K-0)vertical bar = (9.6(-5.1)(+3.5))%, with r(B)* < 15.0% at 90% confidence level.
|
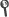
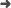
|
|
|
BABAR Collaboration(del Amo Sanchez, P. et al), Lopez-March, N., Martinez-Vidal, F., Milanes, D. A., & Oyanguren, A. (2010). Observation of the Y(1(3)D(J)) bottomonium state through decays to pi(+)pi Y-(1S). Phys. Rev. D, 82(11), 111102–7pp.
Abstract: Based on 122 X 10(6)Y(3S) events collected with the BABAR detector, we have observed the Y(1(3)D(J)) bottomonium state through the Y(3S) -> gamma gamma Y(1(3)D(J)) -> gamma gamma pi(+)pi Y-(1S) decay chain. The significance for the J = 2 member of the Y(1(3)D(J)) triplet is 5.8 standard deviations including systematic uncertainties. The mass of the J = 2 state is determined to be 10 164.5 +/- 0.8(stat) +/- 0.5(syst) MeV/c(2). We use the pi(+)pi(-) invariant mass distribution to confirm the consistency of the observed state with the orbital angular momentum assignment of the Y(1(3)D(J)).
|
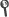
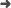
|
|
|
BABAR Collaboration(del Amo Sanchez, P. et al), Lopez-March, N., Martinez-Vidal, F., Milanes, D. A., & Oyanguren, A. (2010). Study of B -> X gamma decays and determination of vertical bar V-td/V-ts vertical bar. Phys. Rev. D, 82(5), 051101–8pp.
Abstract: Using a sample of 471 x 10(6) B (B) over bar events collected with the BABAR detector, we study the sum of seven exclusive final states B -> X-s(d)gamma, where X-s(d) is a strange (nonstrange) hadronic system with a mass of up to 2.0 GeV/c(2). After correcting for unobserved decay modes, we obtain a branching fraction for b -> d gamma of (9.2 +/- 2.0(stat) +/- 2.3(syst) x 10(-6) in this mass range, and a branching fraction for b -> s gamma of (23.0 +/- 0.8(stat) +/- 3.0(syst) x 3.0(syst) x 10(-5) in the same mass range. We find B(b -> d gamma)/B(b -> s gamma) = 0.040 +/- 0.009(stat) +/- 0.010(syst), from which we determine vertical bar Vtd/Vts vertical bar = 0.199 +/- 0.022(stat) +/- 0.024(syst) +/- 0.002(th).
|
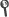
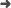
|
|
|
BABAR Collaboration(del Amo Sanchez, P. et al), Lopez-March, N., Martinez-Vidal, F., Milanes, D. A., & Oyanguren, A. (2010). Observation of new resonances decaying to D pi and D*pi in inclusive e(+)e(-) collisions near root s=10.58 GeV. Phys. Rev. D, 82(11), 111101–9pp.
Abstract: We present a study of the D+pi(-), D-0 pi(+), and D*(+)pi(-) systems in inclusive e(+)e(-)-> c (c) over bar interactions in a search for new excited D meson states. We use a data set, consisting of similar to 454 fb(-1), collected at center-of-mass energies near 10.58 GeV by the BABAR detector at the SLAC PEP-II asymmetric-energy collider. We observe, for the first time, candidates for the radial excitations of the D-0, D*(0), and D*(+), as well as the L = 2 excited states of the D-0 and D+, where L is the orbital angular momentum of the quarks.
|
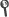
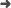
|
|
|
Sanjuan, R., Nebot, M., Chirico, N., Mansky, L. M., & Belshaw, R. (2010). Viral Mutation Rates. J. Virol., 84(19), 9733–9748.

Abstract: Accurate estimates of virus mutation rates are important to understand the evolution of the viruses and to combat them. However, methods of estimation are varied and often complex. Here, we critically review over 40 original studies and establish criteria to facilitate comparative analyses. The mutation rates of 23 viruses are presented as substitutions per nucleotide per cell infection (s/n/c) and corrected for selection bias where necessary, using a new statistical method. The resulting rates range from 10(-8) to 10(-6) s/n/c for DNA viruses and from 10(-6) to 10(-4) s/n/c for RNA viruses. Similar to what has been shown previously for DNA viruses, there appears to be a negative correlation between mutation rate and genome size among RNA viruses, but this result requires further experimental testing. Contrary to some suggestions, the mutation rate of retroviruses is not lower than that of other RNA viruses. We also show that nucleotide substitutions are on average four times more common than insertions/deletions (indels). Finally, we provide estimates of the mutation rate per nucleotide per strand copying, which tends to be lower than that per cell infection because some viruses undergo several rounds of copying per cell, particularly double-stranded DNA viruses. A regularly updated virus mutation rate data set will be available at www.uv.es/rsanjuan/virmut.
|
|

